|
http://www.abbs.info e-mail:[email protected] ISSN 0582-9879 ACTA BIOCHIMICA et BIOPHYSICA SINICA 2002, 34(6): 690-696 CN 31-1300/Q |
Purification
and Characterization of a Novel Chitinase from Bacillus brevis
(
Department of Biochemistry, School of Life Sciences, Fudan University, Shanghai
200433, China;
1Shanghai
Academy of Agricultural Sciences, Shanghai 201106,
China )
Recently
chitin and chitinases are receiving more and more attention from biologists. A
wide variety of medical applications of chitin and chitin derivatives have been
reported over the last three decades[14]. N, N′-diacetylchitobiose
has been widely used as starting material for synthesis of biological active
compounds[15,16]. Chitinases promise to be safer pesticides (than
chemical ones) and microbial biocontrol agents due to the importance of
chitinolytic enzymes in insect, nematode, and fungal growth and development[17].
Chitinase activity in human serum has recently been detected, and it may play a
role in defending the invasion of fungal pathogens[18].
Bacteria,
fungi, plants and insects are four major objects of chitinase research. In
bacteria, Bacillus[19] and Streptomyces[20]
are intensively studied for their high productivity of chitinases. Wiwat et
al.[19]reported that Bacillus circulans WL-12
secreted chitinases into the culture medium, among which chitinase A1 showed
strong affinity to chitin and played a major role in the hydrolysis of chitin.
Chitinases have also been found in other Bacillus species including B.cereus,
B.licheniformis and B.subtilis[21].
Bacillus
brevis No.G1 is a newly isolated strain from
soil in Shanghai of China for its high chitinase activity secreted in culture
medium[22]. In this study we purified the extracellular chitinase to
homogeneity from the fermented broth of B.brevis No.G1 and investigated
its physico-chemical properties. With the determination of partial N-terminal
amino acid sequence and its characteristics, we demonstrated that the purified
enzyme was a novel endochitinase.
1.1
Bacterial strain and culture condition
Bacillus
brevis No.G1 was isolated from soil in Shanghai
of China as previously reported[22]. Cultures were maintained on
nutrient agar slants and incubated at 30 ℃
for 72 h. The bacterial cells were then inoculated into a
500-ml Erlenmeyer flask containing 50 ml liquid medium, cultured at 30 ℃
for 48-72
h on a shaker until most spores broke off. The liquid medium for bacterial
growth contained 20 g/L soybean powder, 4 g/L starch, 3 g/L peptone, 2 g/L
yeast extract, 0.3 g/L KH2PO4, 0.2 g/L MgSO4,
and 1 g/L CaCO3.
1.2
Chemicals
Phenyl-Sepharose
CL-4B, DEAE-Sepharose Fast Flow, Sephadex G-150 were purchased from Pharmacia
LKB (Uppsala Sweden). Purified chitin, chitosan and thin-layer chromatography
(TLC) plates were purchased from Sigma. Other chemicals were of analytical
grade.
1.3
Preparation of colloidal chitin
Colloidal
chitin was prepared from purified chitin according to the method of Roberts et
al.[23] with minor modification. Ten grams of chitin powder were
added slowly into 180 ml of HCl (37%, W/V) at 25 ℃
under vigorous stirring for 2 h. The suspension was poured into 1 liter of
ice-cold 95% alcohol under vigorous stirring for 30 min, and stored at -20 ℃
until use. When in need, 10 ml of the suspension was centrifuged. The
precipitate was washed with 50 ml of 0.1 mol/L sodium phosphate buffer (pH 7.0)
for 3 times. The derived precipitate was dissolved in 90 ml 0.1 mol/L sodium
phosphate buffer (pH 6.0), which was about 10 g/L colloidal chitin solution.
1.4
Purification of chitinase
The
fermented broth of B. brevis No.G1 was collected by brief centrifugation
and the proteins fractionated with 50% saturation (NH4)2SO4
were collected by centrifugation at 8 000 g for 20 min. The protein
precipitate was dissolved in 0.8 mol/L (NH4)2SO4
solution and the insoluble materials were removed by centrifugation at 15 000 g
for 30 min. The derived supernatant was applied onto a Phenyl-Sepharose CL-4B
column (f1.2
cm×10 cm) pre-equilibrated with 1 mol/L (NH4)2SO4.
The column was washed with 1.5 bed volumes of 1 mol/L (NH4)2SO4,
2 bed volumes of 0.1 mol/L (NH4)2SO4, and then
eluted with distilled water. The flow rate was maintained at 0.5 ml/min. The fractions
with chitinase activity were pooled and dialyzed overnight at 4 ℃
against 10 mmol/L Tris-HCl buffer, pH 8.2. The dialysate was collected and
immediately applied on a DEAE-Sepharose Fast Flow column (f1.2
cm×4 cm) pre-equilibrated with 10 mmol/L Tris-HCl
buffer, pH 8.2. The column was further washed with 1.5 bed volumes of 10 mmol/L
Tris-HCl buffer (pH 8.2) and then developed with linear 0.04-0.14
mol/L NaCl gradient in 10 mmol/L Tris-HCl buffer, pH 8.2. The flow rate was
maintained at 0.25 ml/min. The fractions with chitinase activity were pooled,
concentrated and kept at -20 ℃
until use.
For
N-terminal amino acid sequencing, the enzyme fraction was further purified by
reverse-phase HPLC. Chitinase fractions derived from DEAE-Sepharose Fast Flow were
loaded on a Zorbax 300SB-CN column (du Pont, f250
mm×4.6 mm I.D.), and the column was
developed with acetonitrile gradient supplemented with 0.1% trifluoroacetic
acid (TFA). The elution pattern was monitored by absorbance at 220 nm and 280
nm. The major peak was collected and lyophilized for automatic amino acid
sequencing.
1.5
Enzymatic activity assay
Two
methods for enzymatic activity assay were used in this work. For rapidly
tracing chitinase activity during the chromatographic separation processes, 100
ml
of each fraction was mixed with 2 ml 10 g/L chitosan in 50 mmol/L acetate
buffer (pH 5.0) and incubated at 50 ℃
for 10 min. The viscositic changes of the chitosan solutions were checked with
naked eye to determine those fractions with chitinase enzyme activity.
Quantitative assay of chitinase activity was carried out by colorimetric method
described by Imoto et al.[24]. In a typical reaction, 50 ml
enzyme solution, 0.05 ml 10 g/L colloidal chitin and 0.35 ml of 0.1 mol/L
Tris-HCl buffer (pH 8.0) were mixed and incubated at 60 ℃
for 15 min. Reaction was terminated by heating in boiling water for 15 min, and
mixed with 2.0 ml of 1.5 mmol/L potassium ferricyanide solution. The mixture
was then heated in boiling water for another 15 min and cooled to room
temperature. The supernatant was subjected to spectrophotometry measurement at
420 nm. The enzyme activity was calculated from a standard curve obtained with
known concentration of GlcNAc. One unit of chitinase activity was defined as
the amount of enzyme that liberated 1 mmol
GlcNAc per min at pH 8.0 and 60 ℃.
Negative control tubes contained all components except substrate, and blanks
contained all components except the enzyme.
1.6
Electrophoresis
SDS-polyacrylamide
gel electrophoresis (SDS-PAGE) was performed according to the method described
by Sambrook et al.[25]. The proteins were stained with
Coomassie brilliant blue R-250. For isoelectric focusing (IEF) experiment,
about 5 mg
of sample proteins in 20 ml
solution were loaded to a precasted capillary gel with 0.75% Ampholine (pH
range 3.5-10)[26],
and run under 200 V for 5 h.
Amyloglucosidase (pI 3.6), trypsin inhibitor (pI 4.6), b-lactoglubin
A (pI 5.1), conalbumin (pI 6.0), myoglobin (pI 6.8, 7.2), lentil lectin (pI
8.2, 8.6, 8.8), and trypsinogen (pI 9.3) were used as markers.
1.7
Zymogram
To
identify the protein of chitinase, zymographic approach was applied on samples
derived from DEAE Fast Flow chromatography. Samples were separated on 10%
polyacrylamide gel electrophoresis at pH 8.3. As soon as the electrophoresis
was finished, the gel was immediately placed on an agarose slab gel containing
10 g/L colloidal chitin. After incubation for 1.5 h at 37 ℃,
a transparent band could be seen on the agarose slab. The sections of the
polyacrylamide gel overlapping with the transparent band were carefully cut out
and pestled with sample buffer in an Eppendorf tube. The derived paste was
analyzed on 10% SDS-PAGE.
1.8
Gel filtration chromatography
Gel
filtration chromatography was used for determination of molecular weight of
chitinase. Sephadex G-150 was packaged in a 1.2 cm ×60
cm column and pre-equilibrated with 10 mmol/L Tris-HCl buffer (pH 8.2) at a
flow rate of 0.15 ml/min. About 0.4 ml concentrated enzyme obtained from the
DEAE Fast Flow column was applied to the column and eluted with the same
buffer. Protein profile was monitored at 280 nm. The molecular weight was
estimated from a standard curve obtained from the proteins with their molecular
weights known.
1.9
Determination of protein concentration
Protein
concentrations were determined by Folin phenol method[27], with
bovine serum albumin as the reference.
1.10 Silica thin layer chromatography
Silica
thin layer chromatography (TLC) was performed according to the method described
by Xia et al.[28]. Five ml
of chitosan samples treated with HCl or digested with purified chitinase, were
applied on TLC plate, and the plate was developed by n-butanol ∶
acetic acid ∶
water (2 ∶
1 ∶ 1). For chitinase digestion, 1 ml of 1%
chitosan in 50 mmol/L acetate buffer (pH 5.0) was mixed with 6 mg
purified enzyme and incubated at 60 ℃
for 1 h, or at 37 ℃
for 24 h. For HCl treatment, 10 g/L chitosan in 4 mol/L HCl was prepared and
heated at 90 ℃
for 2 h. GlcNAc and chitotetraose were used as markers for TLC.
1.11 N-terminal amino acid sequencing
Samples
obtained from reverse-phase HPLC were lyophilized and subjected to N-terminal
amino acid sequencing on an automatic protein sequencer (Model 473A, Applied
Biosystems Inc, USA)
2.1
Purification of chitinase
B.brevis
No.G1 was originally isolated from soil of
Shanghai urban for its high chitinase activity secreted into the culture media[22].
We purified a chitinase secreted by this new strain of B.brevis for
further study. Proteins in the fermented broth was recovered with (NH4)2SO4
precipitation at 50% saturation. The protein precipitates were dissolved in 0.8
mol/L (NH4)2SO4 solution and separated by
Phenyl-Sepharose CL-4B hydrophobic interaction chromatography. In a typical
separation, 30 ml of the sample solution containing 68.4 mg of crude proteins
was applied to a 1.2 cm×10
cm column, and developed as described in Materials and Methods. Four protein
peaks were detected at 280 nm as shown in Fig.1 and Fig.2. Chitinase activity was
only found in the last peak eluted with distilled H2O.

Fig.1 Elution profile of chitinase on
Phenyl-Sepharose CL-4B column
The protein solution was eluted stepwise
with 0.1 mol/L (NH4)2SO4 and distilled water.
Absorbance at 280 nm (□)
and relative enzymatic activity (●)
were determined.

Fig.2 Elution profile of chitinase on
DEAE-Sepharose Fast Flow column choromatography
The protein solution was eluted with a
linear gradient from 0.04 mol/L NaCl to 0.14 mol/L NaCl in 10 mmol/L Tris-HCl
(pH 8.2). Absorbance at 280 nm (□)
and relative enzymatic activity (●)
were measured.

Fig.3 SDS-PAGE analysis of chitinase under
various conditions
1,5,
protein markers (I: BSA, bovine serum albumin, 68 kD; II: ovalbumin, 45 kD;
III: carbonic anhydrase, 31 kD); 2, chitinase treated with 2-mercaptoethanol;
3, chitinase treated with 0.1% TFA; 4, chitinase heated at 55 ℃
in 8 mol/L urea for 30 min; 6, chitinase recovered from the PAGE gel slice
overlapping the transparent band on zymogram; 7, chitinase derived from DEAE
chromatography.

2.2
N-terminal amino acid sequence
For
N-terminal sequencing, the enzyme fractions from DEAE-Sepharose Fast Flow were
further purified by reverse-phase HPLC. As shown in Fig.4, only one protein
peak was detected. The protein peak was collected, lyophilized and subjected to
amino acid sequencing. The first 10 amino acids in the N-terminal sequence were
determined to be AVSNSKIIGY. The sequence was blasted against GenBank, however,
no chitinase known showed significant similarity with this sequence. The 10
N-terminal amino acids of chitinase A1 from Bacillus circulans
No.4.1 is APWNSKGNYA[19], which was the most close to the N-terminal
amino acid sequence of the B.brevis No.G1 chitinase.

Fig.4 Elution profile of chitinase on
reverse-phase HPLC
The
CN column was developed by a linear gradient of acetonitrile from 35 % to 50%
at the presence of 0.085% TFA.
The
protein in the HPLC peak showed a mass of 48 kD on SDS-PAGE and no enzyme
activity. The difference between molecular weights of proteins in
DEAE-Sepharose Fast Flow fractions and HPLC fractions implied a dimer structure
of chitinase. We performed a series of experiments or further characterization
of this protein. On gel filtration chromatography, the molecular weight of
chitinase was determined to be around 81 kD (Fig.5). The molecular weight of
chitinase protein on SDS-PAGE differed depending on conditions. If the
chitinase in DEAE fractions was heated in boiling sample buffer before SDS-PAGE
analysis, the protein band on SDS-PAGE was at the position of 48 kD; and if not
heated, at the position of 85 kD. After treatment of the enzyme with 4% 2-mercaptoethanol, the molecular weight
of the purified enzyme was still 85 kD on SDS-PAGE, implying that disulfide
bond is not involved in the formation of chitinase dimers. Thus it is a
reasonable conclusion that the chitinase dimer is most likely formed by
hydrophobic interaction. Incubating the chitinase with 8 mol/L urea at 50 ℃
for 30 min could cause a total loss of enzymatic activity and a shift of the
protein band position from 85 kD to 48 kD on SDS-PAGE (Fig.3). After dialysis
of the enzymes depolymerized by 8 mol/L urea, by heated at 100 ℃
or by 0.1% TFA treatment against 10 mmol/L Tris-HCl buffer (pH 8.2), the
enzymatic activity recovered by 79%, 75% and 87%, respectively; and the dimer
was found to be the major component revealed by SDS-PAGE (data not shown).
These results strongly suggest that the chitinase has a homodimer structure
based on hydrophobic interaction. After incubating the chitinase with 8 mol/L
urea and then dialyzing it against 40% alcohol, we obtained free chitinase
subunits, which utterly lost the activity (the chitinase in 40% alcohol still
exhibited hydrolytic activity). It was thus concluded that the compact
structure of the dimer is necessary for chitinase activity.

Fig.5 Determination of molecular weight by
gel filtration
A Sephadex G-150 column (1.2 cm×60
cm) equilibrated with 10 mmol/L Tris-HCl buffer (pH 8.2) was used for
determination of the molecular weight of the purified enzyme. Conalbumin (86
kD), bovine serum albumin (68 kD) and ovalbumin (45 kD) were used as molecular
weight markers. The arrow indicated the position where purified chitinase was
eluted.
The
purified chitinase was subjected to isoelectric focusing analysis and the pI of
the chitinase was found to be 5.5 (data not shown). The optimal conditions for enzymatic reaction were studied
systemically. 6 mg
purified chitinase was used to determine its characteristics. The optimal pH of
the chitinase was 8.0 (Fig.6), which was similar to that of the chitinases from
B.circulans No.4.1[19] and Alteromanas sp.strain O-7[29].
The chitinase was stable and could hydrolyze colloidal chitin at a wide pH range
(from pH 6.0 to pH 10.0). The chitinase exhibited the highest activity at 60 ℃
and retained high activity even over 80 ℃
(Fig.7), but in the absence of substrate the chitinase lost its activity
markedly above 60 ℃
(Fig.8), inferring that the substrate could protect the active center of the
chitinase from denaturation.

Fig.6 Effect of pH on enzymatic activity of
the purified chitinase
The chitinase activity was determined at different
pH at 50 ℃.
Buffers used were 0.1 mol/L sodium acetate buffer (pH 3.0, 4.0, 5.0), 0.1 mol/L
sodium phosphate buffer (pH 6.0, 7.0), 0.1 mol/L Tris-HCl buffer (pH 8.0), 0.1
mol/L glycine-NaOH buffer (pH 9.0, 10.0), 0.1 mol/L Na2HPO4-NaOH
buffer (pH 11.0, 12.0).

Fig.7 Effect of temperature on enzymatic
activity of the purified chitinase
The
enzymatic activity of chitinase was measured at different temperatures in 0.1
mol/L Tris-HCl buffer (pH 8.0).
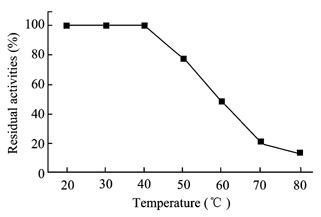
Fig.8 Effect of temperature on stability of
the purified chitinase
The
remained enzymatic activities after incubation at different temperatures in 0.1
mol/L Tris-HCl buffer (pH 8.0) were measured.
2.4
Hydrolysis pattern of the purified chitinase
To
clarify the action mode of the chitinase,viscositic changes of 10 g/L chitosan
solution (in 10mmol/L acetate buffer, pH 5.0) caused by enzymatic hydrolysis
were studied. It was found that the viscosity was promptly reduced in 5 minutes
due to cleavage of chitosan long chains by the chitinase at 60 ℃
(Fig.9). Thus it was concluded that the purified chitinase had endo-splitting
activity.

Fig.9 Hydrolysis of chitosan by the purified chitinase
Purified chitinase (60 mg)
was added to 20 ml 10 g/L chitosan in 50 mmol/L acetate buffer (pH 5.0) and the
mixture was incubated at 60 ℃
for digestion. Aliquots of 2 ml solution were removed at intervals and
subjected immediately to viscosity measurement on an Ostwald viscometer.

Fig.10 TLC analysis of chitosan hydrolysis
1, mixture of 4 mg
GlcNAc and 4 mg
chitotetraose; 2, 10 g/L chitosan in 4 mol/L HCl at 90 ℃
for 2 h; 3, 10 g/L chitosan hydrolyzed by 6 mg
of purified chitinase at 60 ℃
for 1 h; 4, 1% chitosan hydrolyzed by 6 mg
of purified chitinase at 37 ℃
for 24 h.
The
B.brevis No.G1 chitinase
was highly stable (retaining higher than 80% activity) in a wide range of pH
(pH 6.0 to 10.0) and temperature (from 35 ℃
to 72 ℃).
In comparison, the chitinase from B.circulans No.4.1 (from 25 ℃
to 50 ℃)[19]
and Aeromonas sp. 10S-24 (from 30 ℃
to 50 ℃)[30]
exhibit their enzymatic activities in a more narrow temperature range and are
less stable. Furthermore, the purified chitinase from B.brevis No.G1 was
strongly resistant to the hydrolysis by trypsin. A common condition (4 mol/L
urea at 25 ℃)
was not sufficient for trypsin digestion of the chitinase. The chitinase in
fermented broth could be kept at 4 ℃
for at least two months without loss of enzymatic activity. The crude fermented
broth of the B.brevis No.G1 has already been applied to vegetables
against mold diseases with remarkable efficacy (to be published). Therefore, we
expect that the B.brevis No.G1 chitinase could be widely applied as a
new tool of biocontrol.
Acknowledgements The research work was
accomplished in the laboratory of biochemistry and molecular biology of Fudan
University, Shanghai, China. The authors are grateful to Dr.WANG Wei-Rong,
Dr.JIANG Pei-Hong and Dr.QIAN Zhi-Kang for their generous help.
1 Wood WA,Kellogg
ST. Biomass Part B: Lignin, Pectin, and Chitin. Methods Enzymol, 1988, 161:
505-514
2 Wang S,Moyne
A,Thottappilly G, Wu S,
Locy RD, Singh NK. Purification
and characterization of a Bacillus cereus exochitinase. Enzyme
Microb Technol, 2001, 28: 492-498
3 Cohen-Kupiec R,Chet I. The molecular biology of chitin
digestion. Curr Opin Biotechnol,
1998, 9: 270-277
4 Sahai AS, Manocha MS. Chitinases of fungi and plants: Their involvement in morphogenesis and
host-parasite interaction. FEMS Microbiol Rev, 1993, 11:
317-338
5 Turner CM,Adler
PN. The in vitro morphogenesis of Drosophila pupal wings. Mech
Dev, 1995, 52(2-3): 247-256
6 Leah R,Tommerup
H, Svendsen I,Mundy J. Biochemical and molecular characterization of three barley
seed proteins with antifungal properties.
J Biol Chem, 1991, 266: 1564-1573
7 Overdijk B,Van Steijn GJ,Odds FC. Chitinase levels in guinea pig
blood are increased after systemic infection with Aspergillus fumigatus.
Glycobiology, 1996, 6(6): 627-634
8 Gooday GW. Aggressive and defensive
roles for chitinases. EXS, 1999, 87:157-169
9 Svitil AL,Chadhain
SMN, Moore JA,Kirchman DL. Chitin degradation proteins
produced by the marine bacterium Vibrio harveyi growing on
different forms of chitin. Appl Environ Microbiol, 1997, 63: 408-413
10 Wang SL,Chang
WT. Purification and characterization of two bifunctional chitinases/lysozymes
extracellularly produced by Pseudomonas aeruginosa K-187 in a shrimp and
crab shell powder medium. Appl Enviorn Microbiol, 1997, 63: 380-386
11 Patil RS,Ghormade
V,Deshpande MV. Chitinolytic
enzymes: An exploration. Enzyme Microb Technol, 2000, 26: 473-483
12 Deshpande MV. Enzymatic degradation of
chitin and its biological applications. J Sci Ind Res, 1986, 45:
273-281
13 Nielsen MN, Sфrensen
J. Chitinolytic activity of Pseudomonas fluorescens isolates from barley
and sugar beet rhizosphere. FEMS Microbiol Ecol, 1999, 30: 217-227
14 Majeti NV,Kumar
R. A review of chitin and chitosan applications. Reactive & Functional
Polymers, 2000, 46: 1-27
15 Terayama H,Takahashi S,Kuzuhara
H. Large-scale preparation of N,
N′-diacetylchitobiose by enzymic
degradation of chitin and its chemical modifications. J Carbohydr Chem,
1993, 12: 81-93
16 Kobayashi S,Kiyosada T,Shoda
S-i. A novel method for synthesis of chitobiose via enzymatic glycosylation
using a sugar oxazoline as glycosyl donor. Tetrahedron Lett, 1997, 38: 2111-2112
17 Herrera-Estrella A,Chet I. Chitinases in biological control.
EXS, 1999, 87: 171-184
18 Escott GM,Adams
DJ. Chitinase activity in human serum and leukocytes. Infect Immun,
1995, 63(12): 4770-4773
19 Wiwat C,Siwayaprahm
P,Bhumiratana A. Purification
and characterization of chitinase from Bacillus circulans No.4.1. Curr
Microbiol, 1999, 39: 134-140
20 Christodoulou E,Duffner F,Vorgias
CE. Overexpression, purification, and characterization of a thermostable
chitinase (Chi40) from Streptomyces thermoviolaceus OPC-520. Protein
Expr Purif, 2001, 23 (1): 97-105
21 Skerman VBD,Mcgowan V,Sneath
PHA. Approved bacteriol lists of bacterial names. Int J Syst Bacteriol,
1980, 30: 225-420
22 Gu ZR,Ma
CZ,Han CA. Screening and
indentification of chitinase-producing Bacillus spp. and determination
of their chitinase activity. Acta Agriculture Shanghai, 2001, 17(3):
92-99
23 Roberts WK, Selitrennikoff CP. Plant
and bacterial chitinases differ in antifungal activity. J Gen Microbiol,
1998, 134: 169-176
24 Imoto T,Yagishita
K. A simple activity measurement of lysozyme. Agric Biol Chem, 1971, 35:
1154-1156
25 Sambrook J,Fritsch EF,Maniatis
T. Molecular Cloning: A Laboratory Manual,
2nd ed. New York: Cold Spring Habor Laboratory Press, 1989, 882-887
26 Frederick M,Ausubel,
Brent R, Kingston RE, Moore DD,
Seidman JG, Smith JA et al. Short
Protocols in Molecular Biology,
3rd ed. New York: John Wiley & Sons, Inc. 1995
27 Zu J,Cao
KM. Experiments on Biochemistry, Shanghai: Shanghai
Scientific and Technical Publishers, 1981
28 Xia GQ, Jin CS, Yang SJ, Zhang SZ, Jin
C. A novel chitinase having a unique mode of action from Aspergillus fumigatus
YJ-407. Eur J Biochem, 2001, 268: 4079-4085
29 Tsujibo H,
Yoshida Y, Miyamoto K, Imada C,
Okami Y, Inamori Y. Purification, properties, and partial
amino acid sequence of chitinase from a marine Alteromanas sp. strain
O-7. Can J Microbiol, 1992, 38: 891-897
30 Ueda M,Arai
M. Purification and some properties of chitinases from Aeromonas sp. No.
10S-24. Biosci Biotech Biochem, 1992, 56: 460-464
Received:May
17, 2002 Accepted:June
12, 2002
This
work supported by a grant from Shanghai Hengda Scien. & Tech. Dev. Co., Ltd.
*Corresponding
author:Tel/Fax,86-21-55522773; e-mail, [email protected]